CellDesigner 4.0 beta
 CellDesigner 4.0 beta is the new Graphical Notation preview version.
CellDesigner 4.0 beta is the new Graphical Notation preview version.
There are several new features and changes in CellDesigner 4.0 beta. Your feedback on the new functions are most welcome.
* Please note that this is the beta version, newly implemented features may not be fully functional.
* If you create/edit a file with this beta version, your model will not be able to open with the lower version of CellDesigner.
Major New Features:
- Enhanced Graphical Notation
CellDesigner's Graphical Notation conforms to SBGN (Systems Biology Graphical Notation) proposal.such as Gene, Reactions, Protein Residues, Protein / Complex Status.
- Plugin development framework. Develop your plugin on Eclipse, and you can call the plugin from [Plugin] menu.
> Plug-In Tutorial [pdf]
- Macro Functions to provide the easy means to draw the common parts of the model.
- Enhanced KineticLaw Edit Dialog:MathML rendering in KineticLaw.
- Export Species/Reactions information including Notes into CSV file.
- Species ID Edit Dialog
- [Layout] menu: Automatic Layout function (using yFiles layout library.)
- Enhanced Edit Functions
- Layer Function (alpha version)
to add comments / arrows on top of the models.
- Database link to Genome Network Platform

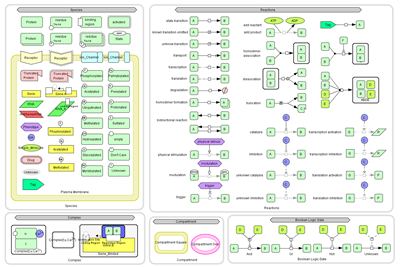
Graphical Notation:
CellDesigner 4.0 beta now enables you to preview for the new graphical notation conform to SBGN Level 1 draft.
(*click to enlarge the upper right image.)
 New Species: Drug, Tag
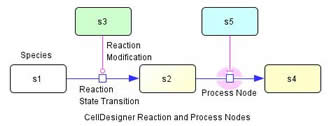
New Species: Drug, Tag - New Reactions: physical stimulation, modulation, trigger, logical gates
- Process nodes of the reactions are introduced to connect modifiers such as catalysis, inhibition.
- Protein states (open, close), binding region

- Protein residue modification types enhanced (Glycosylated, Myristoylated, Palmtoylated, Prenylated, Protonated, Sulfated)
- Gene modification regions enhanced: (phospholyrated, acetylated, methylated),
coding regions, regulatory regions, transcriptionstartingSiteL/R
- Allow to draw Complexes within a Complex
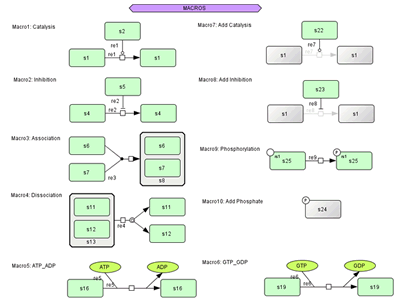
Macro Function:
- 10 quick to draw sets of components available.
KineticLaw Editor: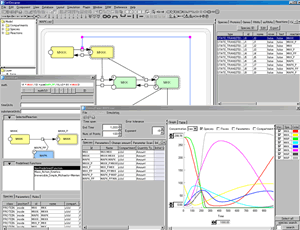
- Preview for the selected reaction in the editor
- Utilize Predefined Function for "math"
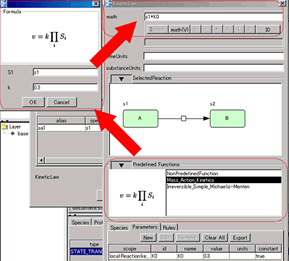
- MathML rendering in "math"
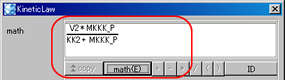
- Switch ID/Name display in the "math"
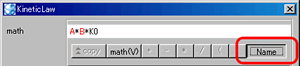
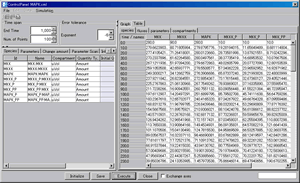 Control Panel:
Control Panel:
- Display simulation results in the tables
- Exchange x-axis and y-axis
File:
- Export Species / Reactions information including Notes into CSV file.
Edit:
- Changes color/shape change for multiple Species/Reactions.
- Highlight List of Proteins selected in the Draw area.
- Sort (ascend/descent) by click in the list
- [Find All] menu added.
- [Species ID Replace] dialog
- Edit the position of the name of the Compartment display
- Reaction ID Display on screen
User Interface Enhancements:
- Change display position of the list: [View]-[List] menu, then [Right] or [Down]
- Display Reaction ID on screen: [View]-[Show Reaction ID]
- Change display position of Compartment name: drag the name to the new position.
- Change Icon display size: [Preference] - [Set Icon Size]
- [Preference]-[Set Macro UI]
- Popup setting dialog: [Preference]-[Popup]
Layer Function (previsionary)
- to add comments / arrows on top of the models.
Installation:
New Installer is being adopted. For details on installation procedure, check here.

 New Species: Drug, Tag
New Species: Drug, Tag 




 Control Panel:
Control Panel: 



